1. Background
Asthma is one of the most common diseases among children with a global prevalence of about 14% and an increasing incidence worldwide (1, 2). It is a disease with heterogeneity and complexity originating from an interplay of gene and environment, persisting and recurring inflammation of the respiratory tract, leading to tissue remodeling and impairing the lung function (3). The pathologic mechanisms of asthma are not completely known yet. Many polymorphisms and cytokines take part in the pathogenesis of inflammation in asthma, such as TGF-β, interleukins (IL-2, IL-4, IL-5, IL-9, IL-6, IL-10, IL-12, IL-13, and IL-17, IL-22, IL-25, IL-33, etc.) (4, 5). TGF-β has played an important role in growth, differentiation, cell migration, formation, and degradation of extracellular matrix components, chemotaxis courses, remodeling, and cell apoptosis in the bronchi. Experimental studies have shown that adenoviral mediated transgenic TGF-β1 in the rodent lung induced serious pulmonary fibrosis and caused the sedimentation of extracellular matrix (6). TGF-β is a multifunctional cytokine, which has both pro-inflammatory and anti-inflammatory effects on airway inflammation and immune response in asthma, and its fibrotic effect has an important role in airway remodeling in asthma (7, 8).
Glucocorticoid (GC) is widely used to treat many chronic conditions, such as asthma, skin diseases, Crohn’s disease, rheumatoid arthritis, and immune rejection after organ transplantation (9, 10). GC binds to its corresponding glucocorticoid receptor (GR) to form the GC-GR complex, that subsequently up-regulates the expression of anti-inflammatory proteins and cytokines (i.e., lipocortin-1, IΚB, MKP1, IL-10, IL-12, and IL-1 receptor antagonist) in the nucleus; meanwhile, it suppresses the expression of pro-inflammatory proteins in the cytoplasm (i.e., IL-2, IL-3, IL-4, IL-5, IL-6, TNF-α, IFN-γ, endothelin-1, and phospholipase A2).
GR is a nuclear receptor superfamily protein, encoded by NR3C1 gene. Studies suggest that mutations and polymorphisms of the NR3C1 gene contribute to a decreased response to GC for the treatment of asthma, leading to drug-resistance (11, 12). Four restriction fragment length polymorphisms (RFLP) (TthIII1, BclI, ER22/23EK, and N363S) may be involved in the pathogenesis of this phenomenon, given their role in the pathology of other diseases, such as metabolic syndrome, autoimmune diseases, and cardiovascular disease (13): TthIII1 coupled with ER22/23EK, is implicated in resistance to GC (11, 14); BclI couples with other two single nucleotide polymorphisms (SNPs, intron 33389 and intron 33388) to increase sensitivity to GC (15, 16); ER22/23EK deceased GC sensitivity (11, 17); the N363S influences the phosphorylation of GR that introduces the structural changes of GR and functional changes of AF1; it is also characterized by increased gene encoding for protein syntheses in response of cells to GC action (17, 18). Over the years, SNPs with altered response to GC therapy have been reported (18-20). TGF-β1 induces the proliferation and chemoattraction of fibroblasts, which results in airway remodeling. It also induces fibroblast to differentiate into myofibroblasts, promoting the production of ECM proteins, fibronectin, and collagen, which finally contributes to the contraction of the ECM (21). GC significantly inhibits the production of TGF-β1 by altering the expression of TGF-β1 mRNA (22, 23), and TGF-β1 induces GC resistance (24). These results imply that SNPs may be associated with TGF-β1, which may illuminate the potential pathological mechanism of children with asthma.
Therefore, we hypothesize that these polymorphisms contribute to the heterogeneity of treatment response given their role in mediating TGF-β1 and GC signaling.
2. Methods
2.1. Subjects
The samples of this study included 52 outpatients (age range: 6 - 14 years) with asthma referred to Huai’an First People’s Hospital, Nanjing Medical University from January 2018 to June 2019. Meanwhile, 40 age-matched healthy volunteers were included as the control group. All the recruited children conformed to the diagnostic guidelines of childhood asthma developed by the national cooperative group on Child Prevention and Treatment in 2016 (25) and The Global Initiative for Asthma (GINA) (2). There were 32 mild, 17 moderate, and three severe cases of asthmatic children who had been diagnosed for 2 - 3 years. The patients had not received any allergen specific immunotherapy, and they had three or more asthma attacks per year; onset was exacerbated at night or early morning. Of the 52 children, 32 were males, and 20 were females. All enrolled children were screened for allergens (serological allergen detection), including 10 non-sensitized, 16 mono-sensitized, and 26 multi-sensitized patients; the total serum of IgE of the allergic patients was above 0.2 IU/L. Meanwhile, 40 healthy children (25 males vs. 15 females) with an age range of 6 - 14 years were recruited and underwent a physical examination. The subjects were included if their immediate relatives across three generations did not have a history of asthma and other lung diseases, such as chronic obstructive pulmonary disease (COPD) and pulmonary fibrosis. There was no history of diabetes, nephritic syndrome, systemic lupus erythematosus (SLE), and psoriasis. Additionally, there was no history of long time use of GC and ephedrine treatment. The study was approved by the ethics committee of Huai’an First People’s Hospital, Nanjing Medical University (Ethical Code: YX-P-2020-002-01). All children or their guardians signed an informed consent.
2.2. Specimen Collection and DNA Extraction
Following the ethical considerations, 200 UL EDTA anticoagulant peripheral fasting blood was collected from both experimental and control groups. The QIAmp DNA Blood Mini Kit (QIAGEN, Germany) was used to extract DNA following manufacturer’s instructions (spin protocol). DNA samples were stored at -20°C.
2.3. Genotyping of TthIII1, BclI, ER22/23EK, N363S: PCR-RFLP Method
The genotyping of TthIII1, BclI, ER22/23EK, and N363S was carried out using primers sequence as previously described (14, 26). The forward primer for TthIII1 was 5’-TCC AGG AGT GGG ACA TAA AGC T-3’, and the reverse primer was 5’-CTT AGA AGC AGA GGT GGA AAT GAA G-3’. The forward primer for BcII was 5'-TGC TGC CTT ATT TGT AAA TTC GT-3', and the reverse primer was 5'-AAG CTT AAC AAT TTT GGC CAT C-3'. The forward primer for ER22/23EK was 5'-GAT TCG GAG TTA ACT AAA AG-3', and the reverse primer was 5'-ATC CCA GGT CAT TTC CCA TC-3'. The forward primer for N363S was 5'-AGT ACC TCT GGA GGA CAG AT-3' and the reverse primer was 5'-GTC CAT TCT TAA GAA ACA GG-3'. Polymerase chain reaction (PCR) was conducted according to the manufacturer’s instructions (Applied Biosystems, USA). The products were digested at 37°C for 4 hours with 1 UL restriction enzyme (Thermo Fisher Scientific, USA). The TthIII1 restriction enzyme, (PsyI) cutting site was 5’…G A C N↓ N N G T C…3’, 3’…C T G N N↑N C A G…5’. The BclI restriction enzyme (BclI) (Thermo Fisher scientific, USA) cutting site was 5’…T↓ G A T C A …3’,3’…A C T A G ↑T…5’. The ER22/23EK restriction enzyme (MnlI) cutting site was 5’…C C T C (N)7 ↓…3’,3’…G G A G (N)6↑ …5’. The N363S restriction enzyme (TasI) cutting site was 5’…↓ A A T T …3’, 3’…T T A A↑T…5’. The RFLP (restriction fragment length polymorphisms) products were isolated using 3% agarose gel electrophoresis, stained with ethidium bromide and observed by UV light. Hydrolyzed fragments of TthIII1 DNA were CT (96bp, 53bp, 43bp), TT (96bp), and CC (53bp, 43bp); DNA fragments of BclI were GC (335bp, 221bp, 117b), CC (221bp, 117bp), and GG (335bp); DNA fragments of ER22/23EK were GG (221bp, 117bp); and DNA fragments of N363S were AA (135bp, 95bp).
2.4. TGF-β1 mRNA Expression
The peripheral mononuclear cells were extracted by Hank’s method from 2 mL EDTA anticoagulant blood. The total RNA was extracted by TRIzol Reagent (Ambion, NY, USA), then 1000 ng RNA was converted to cDNA by reverse transcription using AccuScript PfuUltraII RT-PCR kit (Agilent Technologies, USA). Primers of TGF-β1 of real-time PCR were 5’-GGT ACC TGA ACC CGT GTT GCT-3’ and 5’-TGT TGC TGT ATT TCT GGT ACA GCT C-3’, and the primers of the loading control GAPDH were 5'-AGC CAC ATC GCT CAG ACA-3’ and 5’-GCC CAA TAC GAC CAA ATC C-3’. The amplification of cDNA was done by qRT-PCR kit (Stratagene, USA), using normal two steps: primer annealing temperature was 61°C and annealing time was 40 cycles of 20 seconds. PCR reaction used Agilent Technologies Stratagene Mx3000P. The RT-PCR amplification of the TGF-β1 and GAPDH gene for every sample, GAPDH as internal reference, the CT (threshold cycle) values were analyzed using Mx-Pro software. ΔCT values calculated using the formula: ΔCT = CT, GENE - CT, GAPDH.
2.5. Statistical Analysis
SPSS 26.0 software and GraphPad Prism version 8 were used for statistical analysis. The genotype distribution frequency and allele frequency of each point in each group were calculated and confirmed to be in line with the Hardy-Weinberg equilibrium (P > 0.05). The comparison of alleles and genotypes between groups was performed by analysis of variance (ANOVA), and P < 0.05 indicated a statistically significant difference. Correlation coefficients were analyzed by Spearman’s rank test.
3. Results
Table 1 indicates the frequencies of genotypes and particular alleles of four SNPs. We found no statistically significant differences in TthIII1 and Bcl1 between the asthmatic and control groups (P > 0.05); N363S and ER22/23EK polymorphisms were not detected between the two groups; also, the genotypes of N363S and ER22/23EK were AA and GG, respectively.
| NR3C1 SNP | Asthma Group | Control Group | P |
|---|---|---|---|
| TthIII1 | 0.959 | ||
| Genotype | |||
| TT | 1 (1.92) | 1 (2.50) | |
| CT | 7 (13.46) | 6 (15) | |
| CC | 44 (84.62) | 33 (82.5) | |
| Allele | |||
| C | 95 (91.35) | 72 (90) | |
| T | 9 (8.65) | 8 (10) | |
| BclI | 0.964 | ||
| Genotype | |||
| CC | 2 (3.85) | 2 (5.0) | |
| CG | 16 (30.77) | 12 (30) | |
| GG | 34 (65.38) | 26 (65.0) | |
| Allele | |||
| C | 20 (19.23) | 16 (20) | |
| G | 84 (80.77) | 64 (80) | |
| ER22/23EK | |||
| Genotype | |||
| GG | 52 (100) | 40 (100) | |
| Allele | |||
| A | 0 (0) | 0 (0) | |
| G | 104 (100) | 80 (100) | |
| N363S | |||
| Genotype | |||
| AA | 52 (100) | 40 (100) | |
| Allele | |||
| A | 104 (100) | 80 (100) | |
| G | 0 (0) | 0 (0) |
a Values are expressed as No. (%).
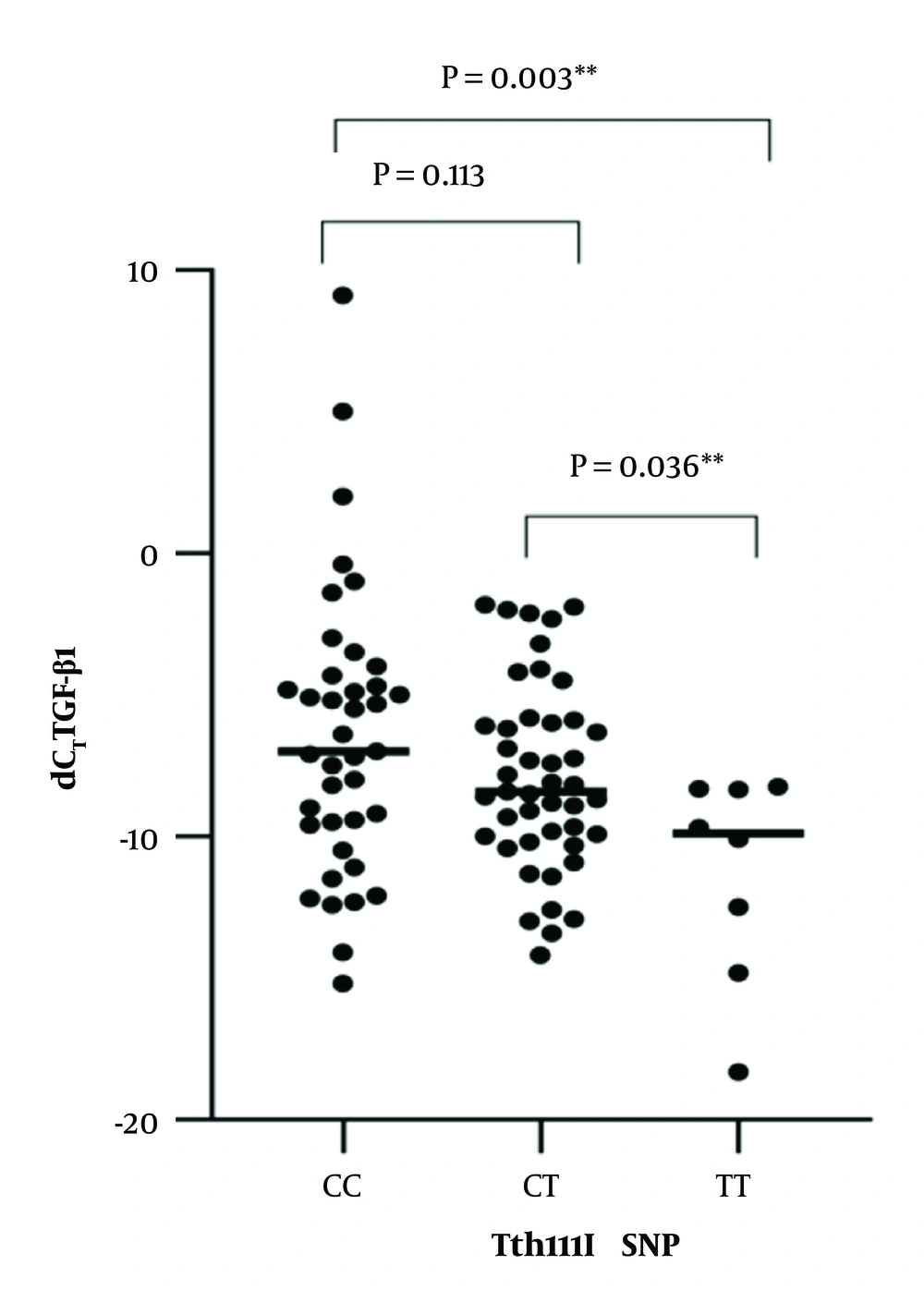
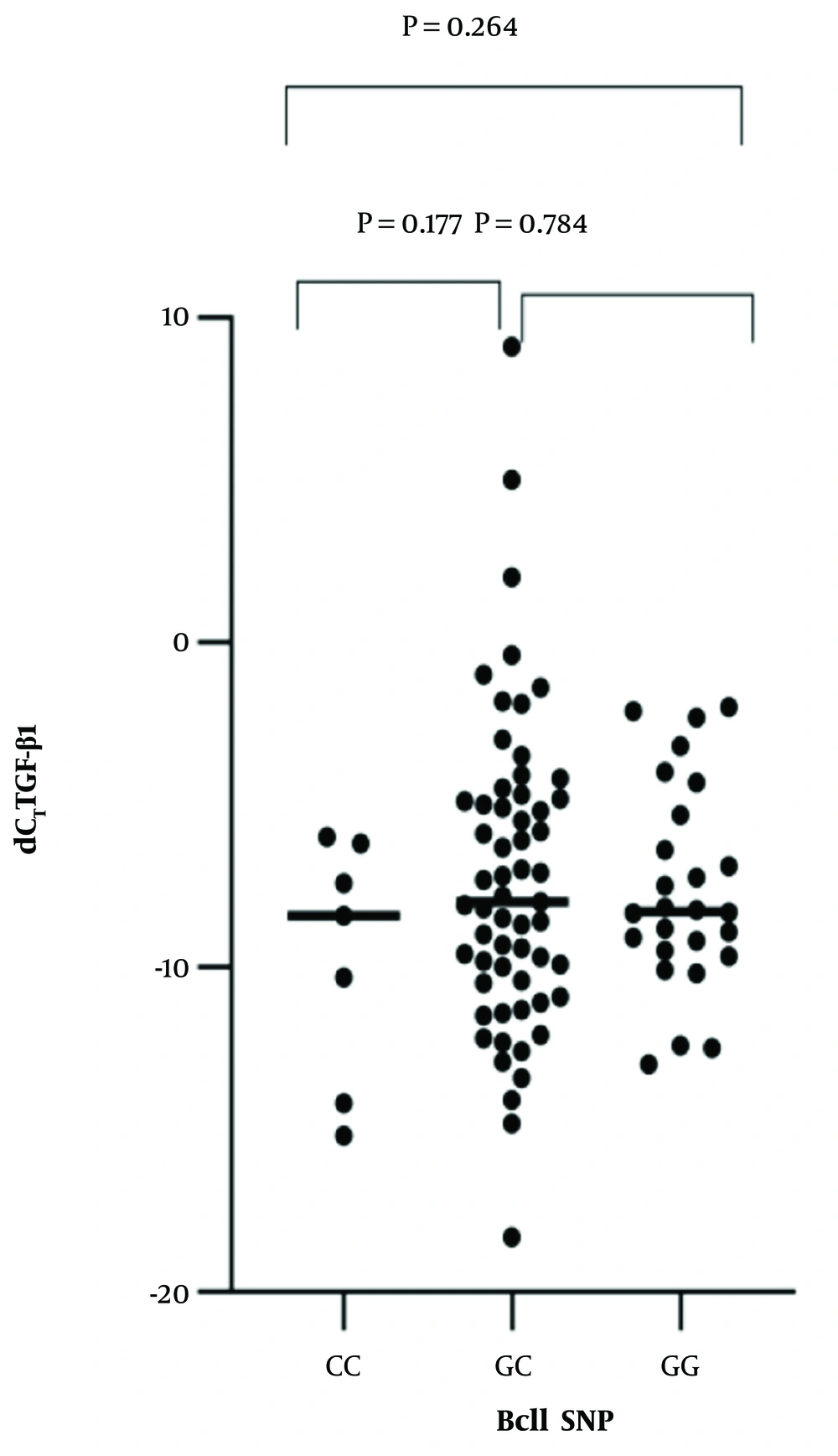
The expression levels of TGF-β1 mRNA are summarized in Table 2. Correlation analysis between the two SNPs and the expression levels of TGF-β1 mRNA were calculated by Spearman’s rank test. TthIII1 and TGF-β1 mRNA were significantly correlated [R2 = -0.25, P = 0.016, beta coefficient = -0.296, 95% CI -0.0147-0.073] (Figure 1). There was no statistically significant correlation between BclI and TGF-β1 mRNA [R2 = 0.063, P = 0.548; beta coefficient = 0.063, 95%CI = 0.036-0.019]. BclI and the level of TGF-β1 mRNA relationships are shown in Figure 2. ER22/23EK and N363S polymorphisms were not found between the groups.
| Parameter | Control Group | Asthmatic Group | Control + Asthmatic Group |
|---|---|---|---|
| Mean ± SD dCT | -9.748 ± 3.268 | -7.459 ± 2.928 | -7.580 ± 4.319 |
| Max. dCT | -4.3 | 9.1 | 9.1 |
| Min. dCT | -18.3 | -14.2 | -18.3 |
4. Discussion
Asthma causes airway hyperresponsiveness, recurrent wheezing, cough, and dyspnea. GC is a vital drug for the treatment of asthma. However, some children with asthma have reduced sensitivity or resistance to it. Studies found that GC resistance is related to the mutation of NR3C1, that changes the amino acid content of the receptor structure and determine the biological action or polymorphism (20, 27-29). Some studies found that glucocorticoid resistant patients have an increased cells expressing IL-2 and IL-4 mRNA compared with numbers seen in steroid-sensitive asthmatic patients in the bronchoalveolar lavage fluid. IL-2 and IL-4 activate P38MAPK kinase, JNK, and ERK, GR phosphorylates at Ser226, which reduces affinity with GC, resulting in reduced sensitivity or resistance to GC (30, 31).
Patients with resistance to GC are associated with epidermal growth factor (EGF). EGF has low expression of phosphorylated tyrosine of these patients, which leading to increasing inflammation and subsequent resistance (32). In asthmatic patients treated with GC, down-regulation of bispecific MAPK phosphatase expression (DUSP1) and loss of MAPK activity affect GC sensitivity (33).
In this study, we found that TthIII1 SNPs are significantly correlated with the expression level of TGF-β1. We hypothesize that TthIII1 SNPs increase the expression level of TGF-β1 and aggravate the inflammation of asthma. TthIII1 CC and CT genotype have the strongest induction effect on the expression of TGF-1. TthIII1 SNPs can significantly increase the expression of TGF-β1 in patients with GC treatment (24); therefore, TGF-β1 can induce and intensify airway remodeling and GC resistance. According to the present study, TthIII1 CC are the most important risk factors for tissue remodeling and GC resistance.
N363 facilitates the action of the GC-GR complex, which improves GC sensitivity (34, 35). It has also been reported that codon N363S AG or GG is related to the enhancement of the anti-inflammatory effect of GC treatment and reduces the risk of out-of-control asthma (28, 29, 34). However, we did not observe this issue in our study, because we did not detect the polymorphism of N363S or racial difference.
Our study found that TthIII1 can induce an increase in the expression level of TGF-β1 mRNA. This causes a progressive asthma course and airway obstruction, leading to the reduction of lung function and poor clinical prognosis. It is thought that TGF-β1 plays a crucial part in the epithelial-mesenchymal transformation (EMT) mechanism, which results in an increase in the number of subepithelial mesenchymal cells, therefore adding to the number of contractile cell and airway hyperreactivity (36). In the course of EMT, epithelial cells lose their classic connections and polarity between cells and acquire a more mesenchymal phenotype. A variety of cytokines, such as interleukins, reinforce MT in respiratory epithelial cells in a TGF-β1 dependent manner, which promotes airway remodeling in patients with asthma. EMT contributes to increasing airway smooth muscle cells in the lung process and elevated TGF-β signaling, this mechanism which has been confirmed by many experiments of animal models of asthma (36).
4.1. Conclusions
The results of this study showed the significant effect of TthIII1 SNPs of NR3C1 polymorphism on increased inflammation in asthmatic children, inducing the expression level of TGF-β1 mRNA. CC and CT of Tth111I are important high-risk factors enhancing the expression level of TGF-β1; therefore, they can result in EMT, contributing to undesirable bronchial remodeling and promotion of airway hyperresponsiveness. The ways to reduce the expression of TGF-β1 and mitigate the degree of airway remodeling have certain implications in the future diagnosis and treatment of bronchial airway asthma. However, the mechanism of bronchial airway remodeling and the interconnection of other growth factors and airway remodeling needs to be further studied. Clinical early intervention in the process of airway remodeling can avoid irreversible airway restriction or inhibit its further development to improve the quality of life of children with asthma.