Introduction
Lung carcinoma represents high metastatic capacity, as well as a high rate of morbidity and mortality. These tumors comprise 17% of all human cancers per year (1). The predominant form of lung cancer is non-small cell lung cancer (NSCLC), accounting for about 85% of all lung cancer cases (2). Metastasis is frequently encountered in NSCLC reported in about 30–40% of the patients (3). Multiple mechanisms and a variety of genetic interactions are involved in this process. In this regard, proto-oncogenes and tumor suppressors are master regulators in cancer pathogenesis and metastasis. Nm23-H1, also known as NDPK-A or NME1, has been identified tumor suppressor gene inhibiting metastasis in various cancers (4). Nm23-H1 has been down-regulated in highly metastatic tumors as mice lacking the Nm23-M1 gene developed lung cancer more frequently than wild-type controls (5). The mechanisms of antitumor activities of Nm23-H1 are yet to be divulged. Nevertheless, modulation of multiple growth factors and matrix metalloproteinases (MMPs) may be involved (6).
The expression of Nm23-H1 is highly regulated by epigenetic factors including histone deacetylases (HDAC). Valproic acid (VPA) is a promising and novel HDAC inhibitor and anti-cancer agent. The administration of VPA has been associated with the downregulation of Nm23-H1 and reduced proliferative capacity of breast cancer cells (7).
Metastasis is a complex phenomenon involving the migration of cells through expressing transmembrane cell adhesion molecules (8). CD44 is a multifunctional transmembrane glycoprotein participating in cell–cell and cell–matrix interactions during tumorigenesis, angiogenesis, tumor growth, and metastasis (9). Particular attention has been given to CD44v6 which is involved in cell–cell and cell–matrix interactions during tumorigenesis (10). Studies have demonstrated the critical role of this protein in the motility and adhesion of cancerous cells to the base membranes (11). It has recently been noted that concomitant inhibition of CD44v6 and matrix metalloproteinase-9 (MMP-9) lowers migration of neoplastic cells (12).
We here assessed the effects of VPA (as a new HDAC inhibitor) on the expression of Nm23-H1 and CD44v6 (as a novel tumor suppressor and a metastasis marker, respectively) in the A549 cell line of NSCLC.
Experimental
Ethics Statement
The study was approved by the Baqiyatallah University of Medical Sciences (code: IR. BMSU.REC.1395.1203).
Cell Culture
The A549 human NSCLC cell line was purchased from Pasture Institute of Iran (Tehran, Iran). The cells were grown in Dulbecco’s modified Eagle’s medium (DMEM /F-12 with GlutaMAX, Gibco, 10565018; Thermo Fisher Scientific Inc., Waltham, MA), which was supplemented with 10% fetal bovine serum (Gemini Bio-Products, West Sacramento, CA), 100 U/mL penicillin G, and 10 mg/mL streptomycin (Invitrogen, Carlsbad, CA). The culture bottles were incubated at 37 °C and 5% CO2. The culture medium was refreshed every 3–4 days. The cells were detached from the old culture using 0.25 mg/mL trypsin/EDTA (Invitrogen).
MTT assay
The viability of the A549 cells treated with VPA was assessed using the standard 3-(4, 5-dimethylthiazol-2-yl)-2, 5-diphenyltetrazolium bromide (MTT) assay. Briefly, the cells were seeded at 104/cells per well in a 96-well plate and incubated overnight at 37 °C and 5% CO2 for 24 h. After refreshing the culture medium, the cells were treated with various concentrations of VPA (0–16 mM) for either 24 h, 48 h, or 72 h. Then, MTT reagent was added at the final concentration of 500 μg/mL to each well, and the plate was further incubated at 37 °C for 4 h in the dark. Finally, the supernatant was discarded and 150 μL DMSO was added to each well. The absorbance was measured at 570 nm with a reference filter of 630 nm using the Synergy H1 Hybrid Multi-Mode Reader (BioTek Instruments, Inc., Winooski, VT). The percentage of alive cells in the presence of VPA was determined respective to the control cells grown in the absence of VPA (13). The IC50 was determined using GraphPad Prism software version 7.03.
Measurement of caspase-3 activity
To measure caspase-3 activity, a caspase-3 substrate (DEVD-pNA, BioVision, Inc., Milpitas, CA) was utilized. For this purpose, the cells treated with various doses of VPA were lysed using chilled cell lysis buffer (BioVision, Inc.). After measuring the protein concentration of the cell lysate using the bicinchoninic acid (BCA) method, equal volumes of protein (100 μg) were diluted to a total volume of 50 μL and mixed with 50 μL of 2X reaction buffer (BioVision, Inc.). The DEVD-pNA substrate was then added to the diluted cell lysates and incubated at 37 °C for 2 h (14). Finally, the absorbance of the released pNA was measured at 405 nm.
RNA extraction and real-time polymerase chain reaction (RT-PCR)
According to the manufacturer’s guidelines, the total RNA was extracted using RNX Plus reagent (Cinnagen, Iran). The quality and quantity of the extracted RNA were assessed by measuring the absorbance ratios of 260/230 nm and 260/280 nm, respectively (NanoDrop spectrophotometer, BioTek, USA). The extracted RNA was further purified using DNase I (Thermo Fisher Scientific Inc.) digestion. Complementary DNA (cDNA) was synthesized using the PrimeScript RT reagent kit (Thermo Fisher Scientific Inc.). Real-time polymerase chain reaction (RT-PCR) was used to determine gene expressions using SYBR Green Master Mix (Parstoos, Iran) and ABI Step One Real-Time PCR System (Applied Biosystems, Foster City, CA). The primers sequences for the genes have been shown in Table 1. All reactions were done in triplicate under the following conditions: 10 min at 95 °C (initial denaturation) followed by 30 cycles as 15 s in 95 °C (denaturation), and 30 s in 60 °C (annealing) and 72 °C (extension). The melting curve analysis was performed within 60 °C to 95 °C. The relative gene expression was calculated using 2–ΔΔCt method (15).
Western blotting
The cells seeded in 6-well plates were scratched and transferred to a microtube. For protein extraction, the cells were suspended in RIPA lysis buffer (Santa Cruz Biotechnology, Inc., Dallas, TX) containing protease inhibitor (Sigma-Aldrich, St. Louis, MO). After that, the cells were sonicated and centrifuged for 10 min at 14,000 rpm and 4 °C. The protein content of the supernatant was evaluated using the BCA method. For each group, 50 µg protein was electrophoresed on 10% SDS-PAGE. The protein bands were then transferred to nitrocellulose membranes (Millipore) using a semi-dry transfer membrane system (Cleaver Scientific Ltd, Warwickshire, United Kingdom). The blocking was performed using 5% skim milk in TBS buffer (20 mM Tris–HCl, 500 mM NaCl, pH 7.4). Mouse anti-human NM23-H1 (Santa Cruz Biotechnology, Inc.) and mouse anti-human CD44v6 (Abcam, Cambridge, UK) antibodies were diluted (1:1000) in TBST buffer (20 mM Tris–HCl, 500 mM NaCl, 0.5% Tween 20, pH 7.4). The membrane was incubated with the primary antibodies overnight at 4 °C. The secondary HRP-conjugated anti-mouse antibody (Santa Cruz Biotechnology, Inc.) diluted in TBST buffer (1:10000) was then added to the membrane and incubated for 1 h at 37 °C. Finally, enhanced chemiluminescence (Thermo Fisher Scientific Inc.) was used followed by exposure to radiographic film to detect secondary antibody binding.
Statistical Analysis
Statistical analysis was performed in SPSS 19. All the tests were done in triplicate. Data were expressed as mean ± SD. One-way ANOVA followed by post hoc Tukey test was used for comparisons between groups. P < 0.05 was considered as the statistical significance threshold.
Results
Cytotoxicity of VPA against A549 Cells
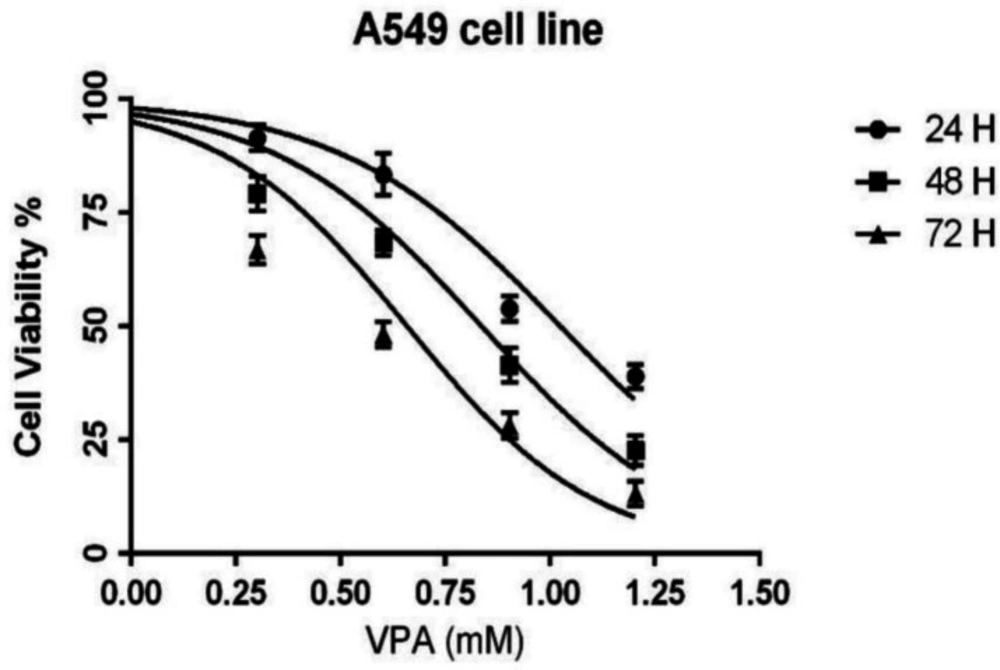
The cells were exposed to different concentrations (0-16 mM) of AVP for either 24, 48 or 72 h. The MTT results showed that VPA inhibited the growth of A549 cells in a concentration- and time-dependent manner (Figure 1). The IC50 values of VPA for A549 cells were 10.5, 6.8 and 4.5 mM in 24, 48 and 72 h incubations, respectively.
VPA promotes caspase-3 activity in A549 cells
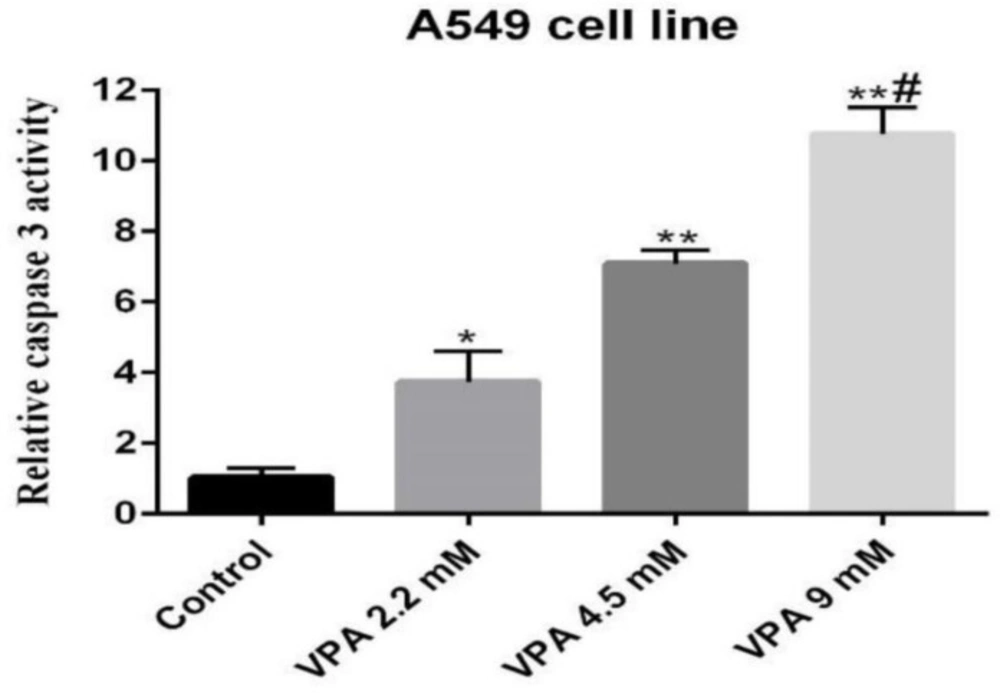
Caspase-3 activity, an early marker of apoptosis, was determined in A549 cells exposed to VPA. Accordingly, the cells exposed to 9 mM of VPA for 72 h showed significantly higher caspase-3 activity respective to other concentrations and periods (P ˂ 0.001, Figure 2).
VPA suppressed MMP-2 and MMP-9 genes expression
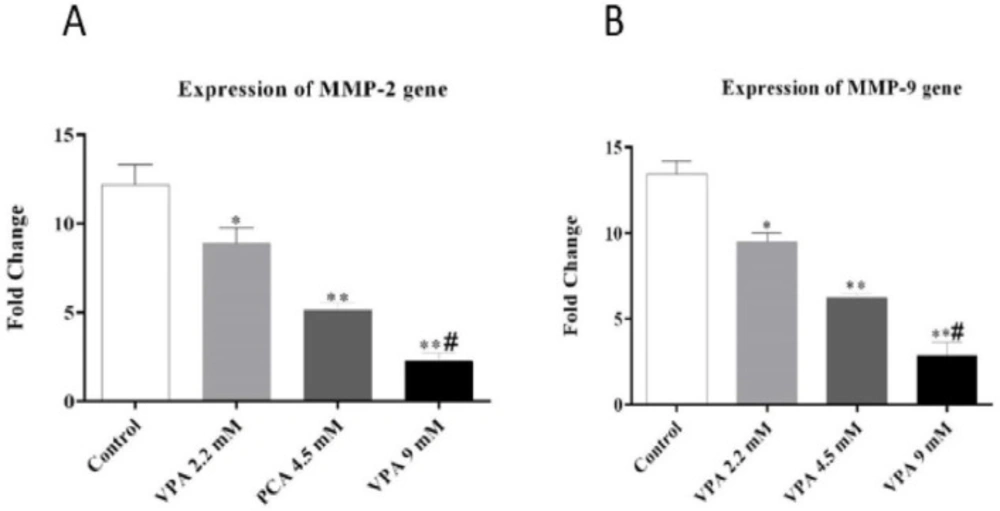
As shown in Figures 3A and 3B, VPA significantly suppressed the expression of MMP-2 and MMP-9 in a dose-dependent manner (P ˂ 0.01 and P ˂ 0.001).
VPA promoted Nm23H-1 and alleviated CD44v6 expression in A549 cells
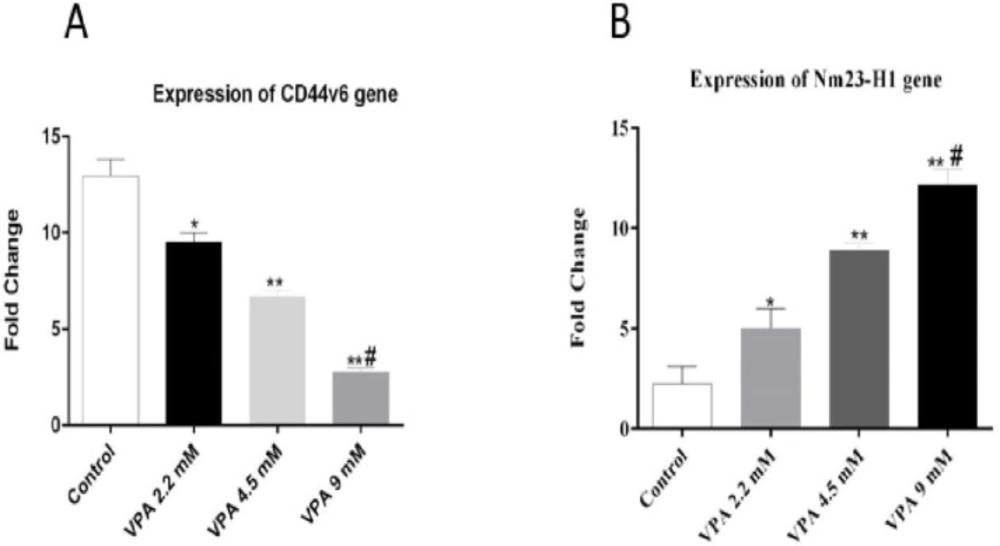
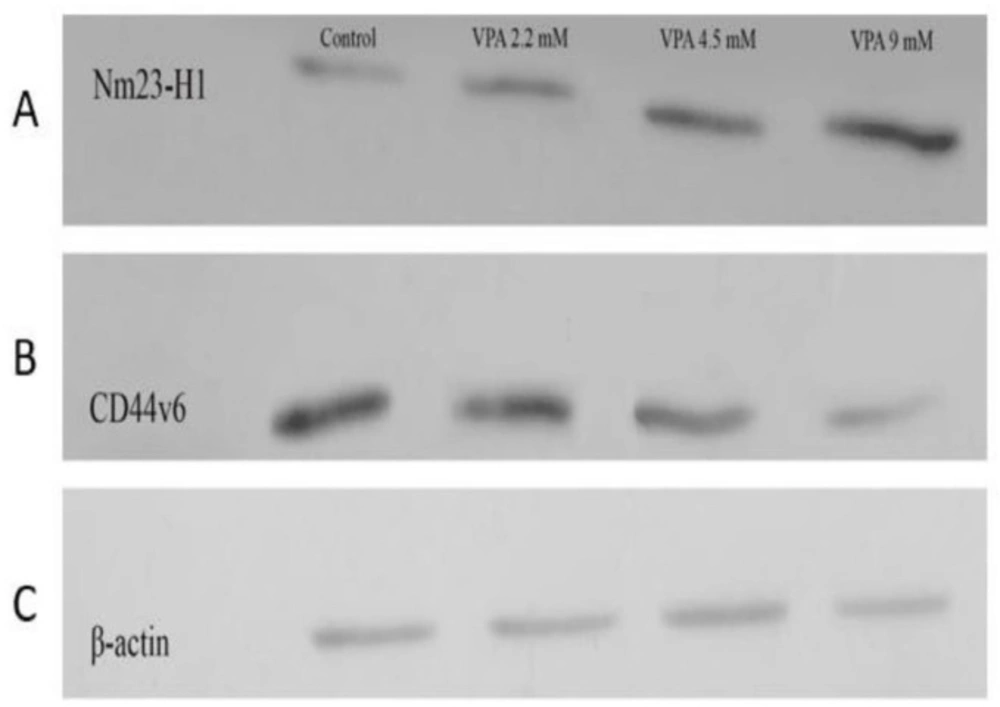
The expressions of Nm23-H1 and CD44v6 andtheir protein levels, were investigated in A549 cells treated with VPA for 72 h. VPA treatment significantly alleviated the expression of CD44v6 gene (Figure 4A, P < 0.001) and protein (Figure 5A) in a dose-dependent way (P < 0.001). On the other hand, the cells exposed to 9 mM VPA represented significantly upregulated Nm23-H1 gene (Figure 4B, P < 0.001) and protein (Figure 5B).
Discussion
In this study, we assessed the effects of VPA, a histone deacetylase inhibitor, to determine its effects on the expression of genes involved in tumor metastasis in the A549 lung cancer cell line. Histone acetylation is one of the most important epigenetic mechanisms modulating gene expression. We observed here that VPA decreased the viability of A549 cells upon 72 h incubation in a dose-dependent manner. Inline, a dose-dependent increase was observed at both gene and protein levels of Nm23-H1, a tumor suppressor in the cancerous cells. On the other hand, the expression of metastatic tumor indicators, CD44v6, MMP-2, and MMP-9 decreased in A549 cells exposed to VPA. These changes further were accompanied by increased caspase-3 activity in VPA treated A549 cells.
Nm23-H1 is a multifunctional protein with nucleoside diphosphate kinase, histidine kinase, and DNase activities (16). Studies have shown the antineoplastic activity of HDACs in hematologic and solid malignancies. Scientific evidence also supports the key role of HDACs in down-regulating of genes involved in tumor metastasis and invasion both in-vitro and in-vivo (17, 18). HDAC inhibitors can suppress tumorigenesis by halting cancer cells migration, invasion, and growth, as well as by inducing apoptosis (19, 20). In this study, we showed that VPA as an HDAC inhibitor, induced the expression of Nm23-H1, a tumor suppressor downregulated in highly metastatic cancers (4). Nm23-H1 upregulation can induce DNA damage and subsequently genomic instability (21). Based on our findings and the effects of VPA on Nm23-H1, epigenetic alternations of this gene in A459 cancer cells may also be a mechanism involved in the metastatic behavior of lung cancer cells. Therefore, applying HDACs inhibitors, particularly VPA, may provide a therapeutic option in this type of cancer.
HDACs also can induce apoptosis in cancer cells (19, 20). Accordingly, it has been shown that VPA-induced apoptosis may involve an increase in acetylation of histones and tubulin, a study conducted in a gastric cancer cell line (22). In our study, the apoptotic effect of VPA on the lung cancer cell line A549 was accompanied by an increase in caspase-3 activity. This result agreed with a previous study that reported the up-regulatory effect of VPA on caspase-3 (23).
CD44 is a cell-surface glycoprotein involved in cell–cell and cell–matrix adhesion, cell migration, and metastasis. Among all CD44 isoforms, CD44v6 harboring a mutation in exon 11 plays an important role in enhancing the adhesive ability of tumor cells (24). The adhesion of cancer cells to the basal epidermal components such as collagen, integrin, and fibronectin is mediated by CD44v6. An evidence-based report showed that an increase in the expression of CD44v6 altered the physicochemical properties of tumor cells and increased their metastatic potential (25). CD44v6 the We showed the inhibitory effects of VPA on CD44v6 at both gene and protein levels in the A459 cell line in the present study. This observation indicates a potential inhibitory impact for VPA on tumorigenesis in lung cancer.
Our results also revealed an inhibitory effect for VPA on the expression of MMP-2 and MMP-9 genes in the A459 cell. Type I collagenases such as MMP-2 and MMP-9 participate in cancer growth and invasion by degrading extracellular matrix (ECM) (26). MMP-2 and MMP-9 are zinc-dependent ECM degrading enzymes involved in the metastatic activity of tumor cells (27). According to the effects of VPA on the expression of these genes in A459 cells observed here, HDACs and epigenetic mechanisms may be involved in lung cancer progression. Therefore, HDACs inhibitors such as VPA can provide a viable therapeutic agent in these cancers.
In the present study, the up-regulation of Nm23-H1 upregulation was seen in concomitant with the down-regulation of MMP-9. This phenomenon can promote a potent anti-metastatic effect on cancer cells. The interaction between Nm23-H1 and MMP-9 is controversial. Nm23-H1 has been shown to increase MMP-9 gene expression and its gelatinolytic activity (28). In another report, however, Nm23-H1 did not modify MMP-9 expression (29). Moreover, Nm23-H1 upregulation has been related to MMP-9 suppression in yet another report (30). In our study, increased Nm23-H1 expression was seen in parallel to decreased expression of CD44v6, MMP-2, and MMP-9. This event, along with elevated caspase-3 activity, can finally result in apoptosis in A549 human lung cancer cells.
In conclusion, our study demonstrated the antitumor activity of VPA against the A549 lung cancer cell line. One possible mechanism may be the Inhibition of HDAC, which increased Nm23-H1 expression. Furthermore, VPA-treated A549 cells showed decreased expression of CD44v6, MMP-2, and MMP-9 considered as metastasis indicators in cancer. The underlying inhibitory mechanisms of VPA on tumor cells and its potential therapeutic role in cancer are yet to be investigated.
| Gene | F/R | Primer sequences (5'3') |
|---|---|---|
| Nm23H1 | Forward | TTAATCAGATGGTCGGGGAT |
| Reverse | GATCTATGAATGACAGGAGG | |
| CD44V6 | Forward | GTCGATGCTAGCTAGCCGTAGCATG |
| Reverse | CGAGCTAGTCGTAGTCGATCGATCG | |
| MMP2 | Forward | TCTCCTGACATTGACCTTGGC |
| Reverse | CAAGGTGCTGGCTGAGTAGATC | |
| MMP9 | Forward | CCTTGTGCTCTTCCCTGGAG |
| Reverse | GGCCCCAGAGATTTCGACTC | |
| GAPDH | Forward | AATCCCATCACCATCTTCCA |
| Reverse | TGGACTCCACGACGTACTCA |